Inputs, outputs, and controls
User inputs, outputs, and control elements on the NGXT site have been redesigned to provide a consistent user experience across the various tools and to present the user with only the most critical decisions and data points. Some of the common elements are described in this section.
Sequence input
Many of the tools on the site can take sequence data, such as proteins or peptides, as input. A typical sequence input box is shown below.
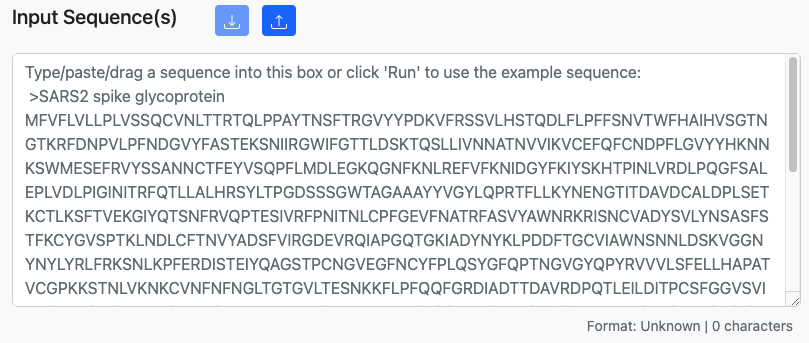
The input box will accept sequences in several common formats:
FASTA
FASTA is a widely used, loosely defined standard for representing amino acid and nucleotide sequences. It generally consists of a descriptor line beginning with a ‘>’ and containing a sequence title, followed by 1 or more lines of amino acid/nucleotide sequences.
Example:
>seq1
LKCFGNTAVAKCNVNHDAEFCDMLRLIDYNKAALSKFKEDVESALHLFKTTVNSLISDQ
LLMRNHLRDLMGVPYCNYSKFWYLEHAKTGETSVPKCWLVTNGSYLNETHFSDQIEQEA
DNMITEMLRKDYIKRQGSTPLALMDLLMFSTSAYLVSIFLHLVKIPTHRHIKGGSCPKP
HRLTNKGICSCGAFKVPGVKTVWKRR
>seq2
YGLKGPDIYKGVYQFKSVEFDMSHLNLTMPNACSANNSHHYISMGTSGLELTFTNDSII
SHNFCNLTSAFNKKTFDHTLMSIVSSLHLSIRGNSNYKAVSCDFNNGITIQYNLTFSDA
QSAQSQCRTFRGRVLDMFRTAFGGKYMRSGWGWTGSDGKTTWCSQTSYQYLII
Whitespace separated
Whitespace separated format is generally ‘lists’ of sequences, without a description, separated by newline characters. This type of input will generally be used for peptide sequences as opposed to full-length proteins, but the latter is also accepted. Note that the input box will treat any whitespace (including spaces, tabs, and newlines) as a sequence delimiter. The following examples are equivalent.
Newline-separated:
LKCFGNTAVA
PYCNYSKFWY
TLMSIVSSLH
Space-separated:
LKCFGNTAVA PYCNYSKFWY TLMSIVSSLH
Named whitespace-separated
This format will have two columns, separated by either spaces or tabs. The first column is a sequence description/title and the second is the peptide/protein sequence.
Example:
seq1 LKCFGNTAVA
seq2 PYCNYSKFWY
seq3 TLMSIVSSLH
The sequence description (column 1) must contain only alphanumeric characters, hyphens (-), underscores (_), or periods (.).
JSON
The JSON format supported by the NGT is a list of objects with a name and a sequence key.
Example:
[
{"name":"seq1",
"sequence":"LKCFGNTAVA"},
{"name":"seq2",
"sequence":"PYCNYSKFWY"},
{"name":"seq3",
"sequence":"TLMSIVSSLH"}
]
Prediction parameters
Default values for most parameters are pre-set in the NGT. However, the user has the option to change the parameters to suit their analysis goals. Most of the controls should be familiar with web components. This section describes those controls that might not be encountered in other contexts.
Prediction models
Specifically part of the T cell - class I application, a variety of different algorithms can be run against the same input dataset. These algorithms are selected in the Prediction Models section of the page. When the page is first loaded the default will be selected, which will look similar to:
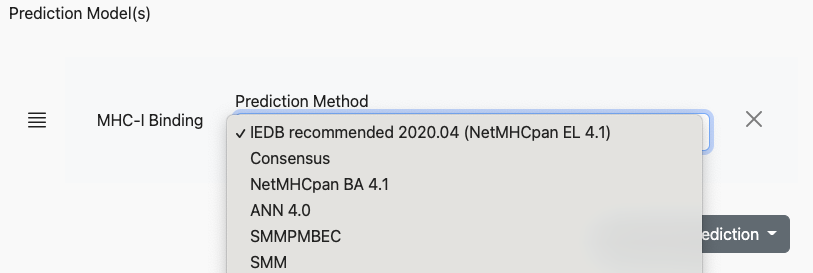
In this configuration, the tool would run an MHC-binding prediction using NetMHCPan EL 4.1 against the input dataset. However, the user may add multiple prediction models to the run, using the Add Another Prediction dropdown:
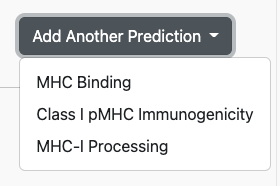
After making a selection here, e.g. MHC-I Immunogenicity, another row of controls will appear with context-specific elements. For MHC-I Immunogenicity, this is a dropdown selector for the Positions to mask:
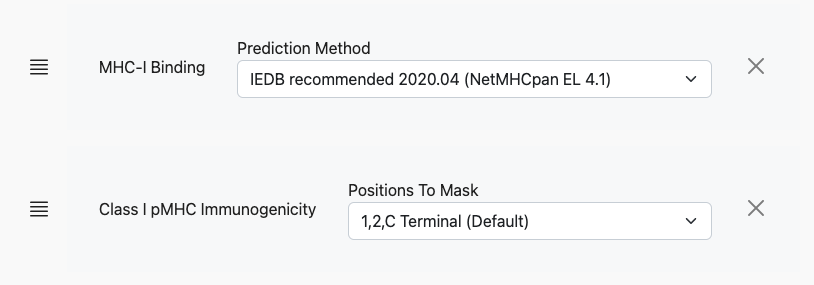
For other predictors, the controls will look different. The user can continue to add prediction models, including additional binding algorithms:
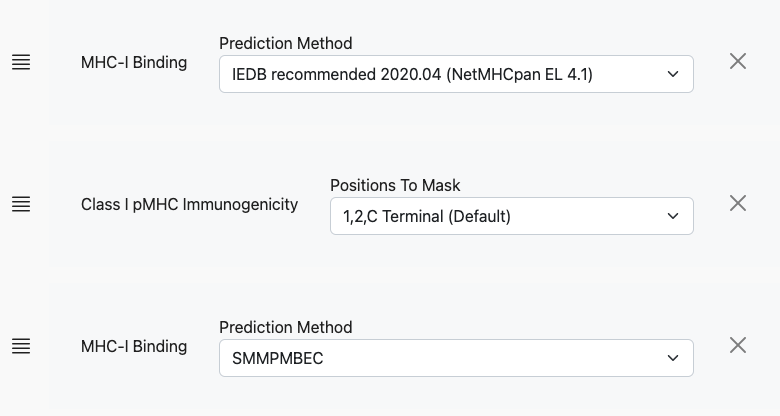
After the tool executes, results will be in tabular format with 1 row per peptide and allele and columns pertaining to each of the prediction models.
Method ordering
Methods may be selected in any order and rearranged by clicking and dragging the ‘handle’ on the left of each set of controls. The only effect that the ordering has is on the ordering of columns in the results table. The columns in the results table will be in the same order as specified by the prediction models.
Running a prediction
Clicking the Run button on an application page will start the prediction with the given inputs and parameters. Depending on the complexity of the analysis, size of the inputs, server load, and other factors a given prediction job can take anywhere from several seconds to hours.
Returning to an analysis
While the system is processing your job, the blue Run button will change to a red Cancel Run button and a loading message will appear beneath it:
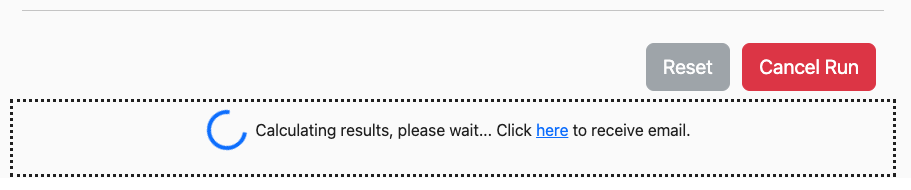
The job will continue to run even if you close your browser window. If you would like to be notified when the job is complete, you can click on the link in the box to provide your email address. Once the job is complete, you will be sent a link back to the page.
You may also return to this page by saving the link from the browser address bar and reloading the page. Each link will contain a unique pipeline ID that will allow you to return to your analysis.
Warnings and errors
Upon submission of your prediction request, the inputs and parameters are evaluated for compatibility.
If issues are discovered that would prevent a portion of your prediction request from running through, these would appear as warnings immediately below the Run button:
The portions of your prediction that are possible will continue to run on the server. However, you may use the Cancel Job button to cancel this request and resubmit after you address the issues with your input and parameter selection.
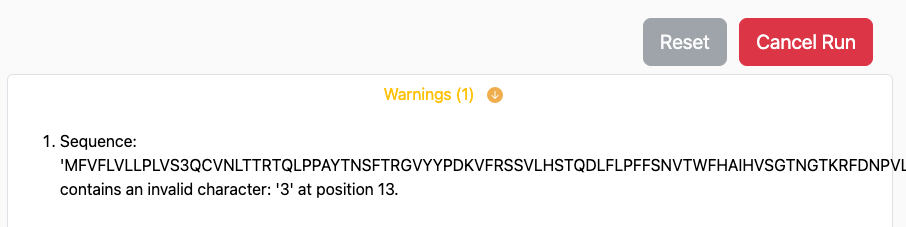
If issues are discovered that would completely prevent the prediction from running through successfully an error is raised and will appear immediately below the Run button, similar to:
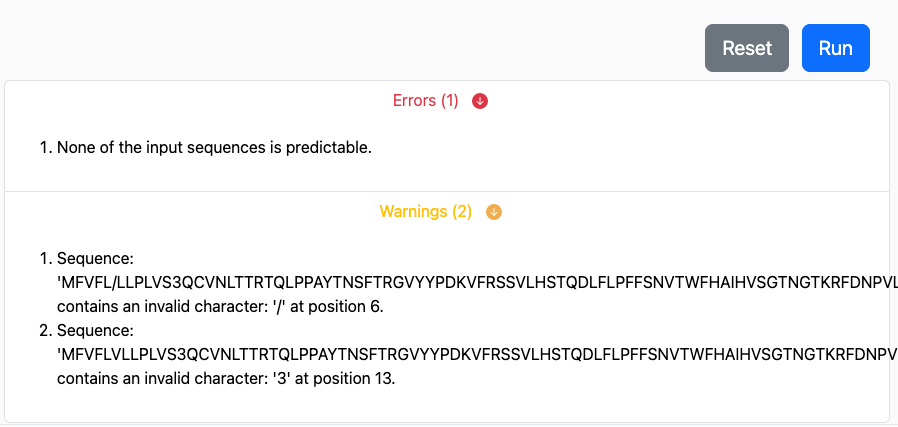
Most warnings and errors will appear almost immediately after submission before any predictions are processed. We aim to make errors and warnings as easily understandable and actionable as possible. Let us know, by sending an email to help@iedb.org if you need assistance interpreting their meaning.
Changing parameters after a prediction
After a prediction has been submitted and results have been returned, it’s possible to change inputs and/or parameters and rerun the prediction. However, in doing so, you will note that any inputs or parameters that have changed are emphasized with a yellow border. For example, if a prediction request is completed for HLA-A02:01 and the user now adds HLA-A*01:01, the allele input control will look similar to below:
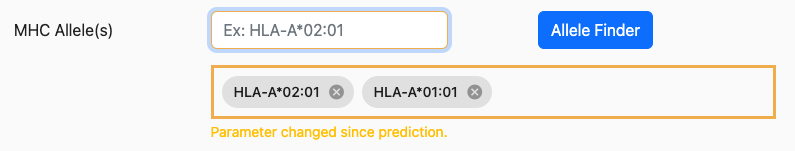
The purpose of this emphasis is to alert users that the results that are currently displayed do not match the currently selected parameters/inputs.
Tabular results
The applications on the NGT may have several types of outputs. The most common type of output will be tabular. Result tables contain a fair amount of functionality that should allow for an elementary analysis of the output. Some of the table functionality and its controls are described in this section.
Sorting & filtering rows
Sorting
Tabular results returned to the user will have a default sorting order defined by the application. This can be overridden by using the controls described below.
Each table header has a set of controls for sorting (up and down arrows) and filtering (filter icon). To sort a table by a specific column, click on either of the arrows. The up arrow will sort in an ascending order while the down arrow will sort in descending order. For instance, to sort on netmhcpan_el_score in descending order, the down arrow in the header would be clicked resulting in:
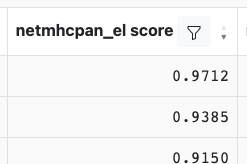
It is also possible to multi-sort, i.e. apply a sorting strategy based on the values in multiple columns. To achieve this, click first on the arrow in the column header to be used for the primary sort. Next, hold down the Shift button on your keyboard while you click on the arrows in the additional column headers that you would like to use for sorting. As you do this, you will notice numbers appearing in the column headers that indicate the order of the column header in the sorting strategy. For instance, if you were to sort first by netmhcpan_el_score, next by smmpmbec_percentile, and finally by immunogenicity_score, the table header would look similar to:

The sort order can be reset back to the default by clicking on the ‘Reset Table’ icon.
Filtering
Clicking on the filter icon in a column header will bring up a menu, similar to:
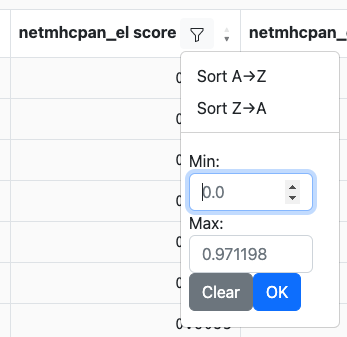
Depending on the data type in the column (e.g., string, integer, etc.) the controls may look slightly different. For instance, numeric columns will allow filtering by value ranges while string columns may allow filtering on specific values. Sticking with the example from above, we can set the minimum score to 0.8:
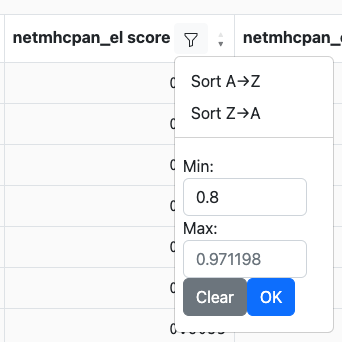
After we click OK, any rows with values in this column that are less than 0.8 will be removed:
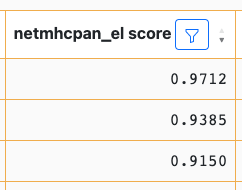

You will note that the cell borders have changed to yellow. This indicates an uncommitted state change of the table, which is described below.
Table states
The state of each analysis, including all parameters, inputs, and the state of the result tables, is tracked by the NGT servers. Applying row filters and changing the columns that are displayed are the two ways that the state of a table can change. Unless this state change is communicated back to the NGT servers, the next time the user loads the analysis from the URL in the address bar the changes to the table will be lost.
As a visual cue to the user, these ‘uncommitted’ state changes are emphasized with yellow borders around each cell of the table. In order to ‘commit’ these changes, click on the ‘Save Table State’ button at the top of the table:

Once this button is clicked, the borders will return to their original color. More importantly, the next time the URL is loaded, the current filters will be applied to the table.
Show/hide columns
Some predictor selections can result in a large number of columns being returned to the user. Most are displayed, but several are kept hidden by default. To change the columns that are displayed, click on the ‘Display Columns’ button at the top of the table:

This will bring up a modal that will list all available columns and allow for the selection/deselection of what is currently displayed:
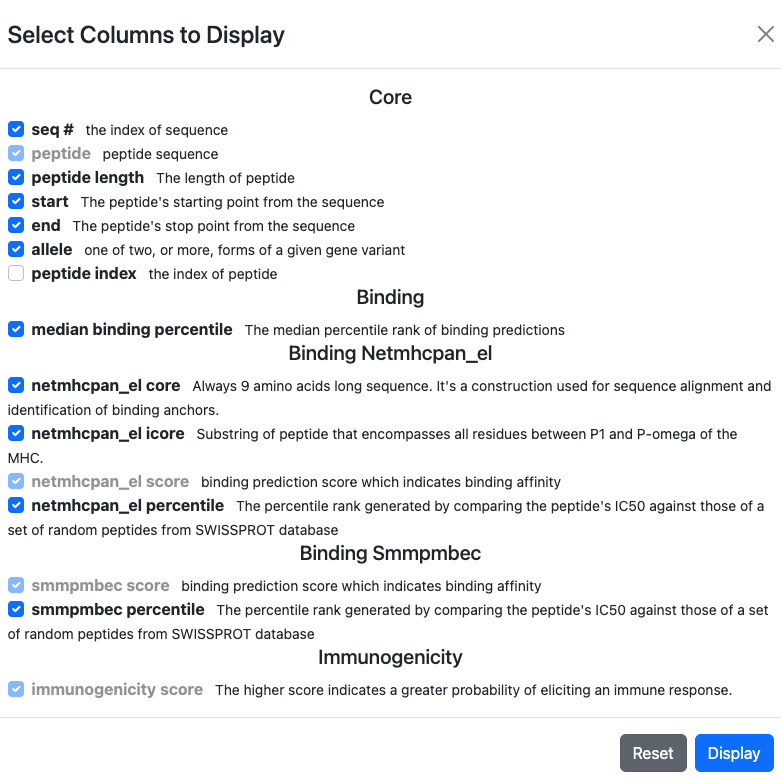
In addition to the name of each column, a brief description of its contents can also be found.
Note that columns that are used for the currently active sort or filter cannot be deselected for display.
To reset the display to the initial defaults, click on the ‘Default’ button.
Downloads
Results can be downloaded in TSV, CSV, and JSON formats by clicking on the ‘Download’ button at the top of the table. A menu will appear that will allow for the selection of ‘All Rows’ versus ‘Displayed Rows’:
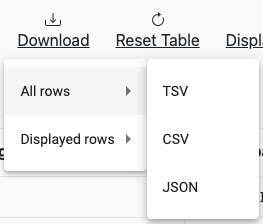
Selecting ‘Displayed Rows’ will download the data after filters are applied.