Mutated Peptide Generator
The Mutated Peptide Generator tool takes a SNPEff-annotated VCF as input and generates predicted neo-peptides and the reference/WT peptides with which they pair.
Methods
MPG reads SNPEff-annotated variants from the VCF, derives mutant and reference amino acid sequences, and generates overlapping neo-peptide and wild-type pairs with configurable length, mutation position(s), frameshift overlap rules, and reference genome build. Optional in-tool SNPEff annotation is supported; pre-annotated VCFs are recommended.
Input Data
The tool accepts VCF files as input. The following steps are recommended before running the tool.
The last line of the VCF headers should look similar to:
#CHROM POS ID REF ALT QUAL FILTER INFO FORMAT {SAMPLE1} {SAMPLE2} ... {SAMPLEN}
The ‘SAMPLE’ columns are not required.
Recommended: Strip Unnecessary Annotations
Stripping unnecessary annotations from the VCF is not required, but recommended as it will help to ensure a consistent
starting point. Annotations can be stripped with vcftools. For example:
vcftools \
--gzvcf infile.vcf.gz \
--recode \
--stdout \
| bgzip -c > infile.stripped.vcf.gz
# index the VCF
tabix infile.stripped.vcf.gz
Recommended: Decomposition and Normalization
Normalization and decomposition of the VCF are highly recommended, but not required. VCF decomposition implies splitting multiallelic variants so each variant has its own record, and normalization generates the most parsimonious allele, meaning that each variant is represented in as few nucleotides as possible.
This step should be done before annotation with SNPEff, using vt. Here is an example command:
zcat < infile.stripped.vcf.gz \
| vt decompose -s - \
| vt normalize -r $REFERENCE - \
| bgzip -c infile.stripped.decomposed.normalized.vcf.gz
# index the VCF
tabix infile.stripped.decomposed.normalized.vcf.gz
The above steps can be combined into one, e.g.:
vcftools \
--gzvcf infile.vcf.gz \
--recode \
--stdout \
| vt decompose -s - \
| vt normalize -r $REFERENCE - \
| bgzip -c infile.stripped.decomposed.normalized.vcf.gz
# index the VCF
tabix infile.stripped.decomposed.normalized.vcf.gz
REFERENCE should point to the genomic fasta file, used for sequence alignments. For instance, Homo_sapiens.GRCh38.dna.primary_assembly.fa.
Recommended: SNPEff annotation
Although we provide the option to run SNPEff annotation on unannotated VCFs, we recommend that users run this before uploading VCFs to the tool.
Note
Other annotators are available, but the MPG tool will only work with SNPEff annotations.
In order to run SNPEff, you will need to prepare a small file that includes the name of the ‘normal’ and ‘tumor’ samples as they appear in the VCF file. In the command below, this is our ‘in.samples’ file. Here is the recommended command, for annotation with GRCh38.86 reference genome:
echo -e "normal_sample\ttumor_sample" > in.samples
zcat < infile.stripped.decomposed.normalized.vcf.gz \
| java -Xmx16G -jar snpEff.jar -cancer -cancerSamples in.samples GRCh38.86 \
| > infile.ann.vcf
Parameter Selection
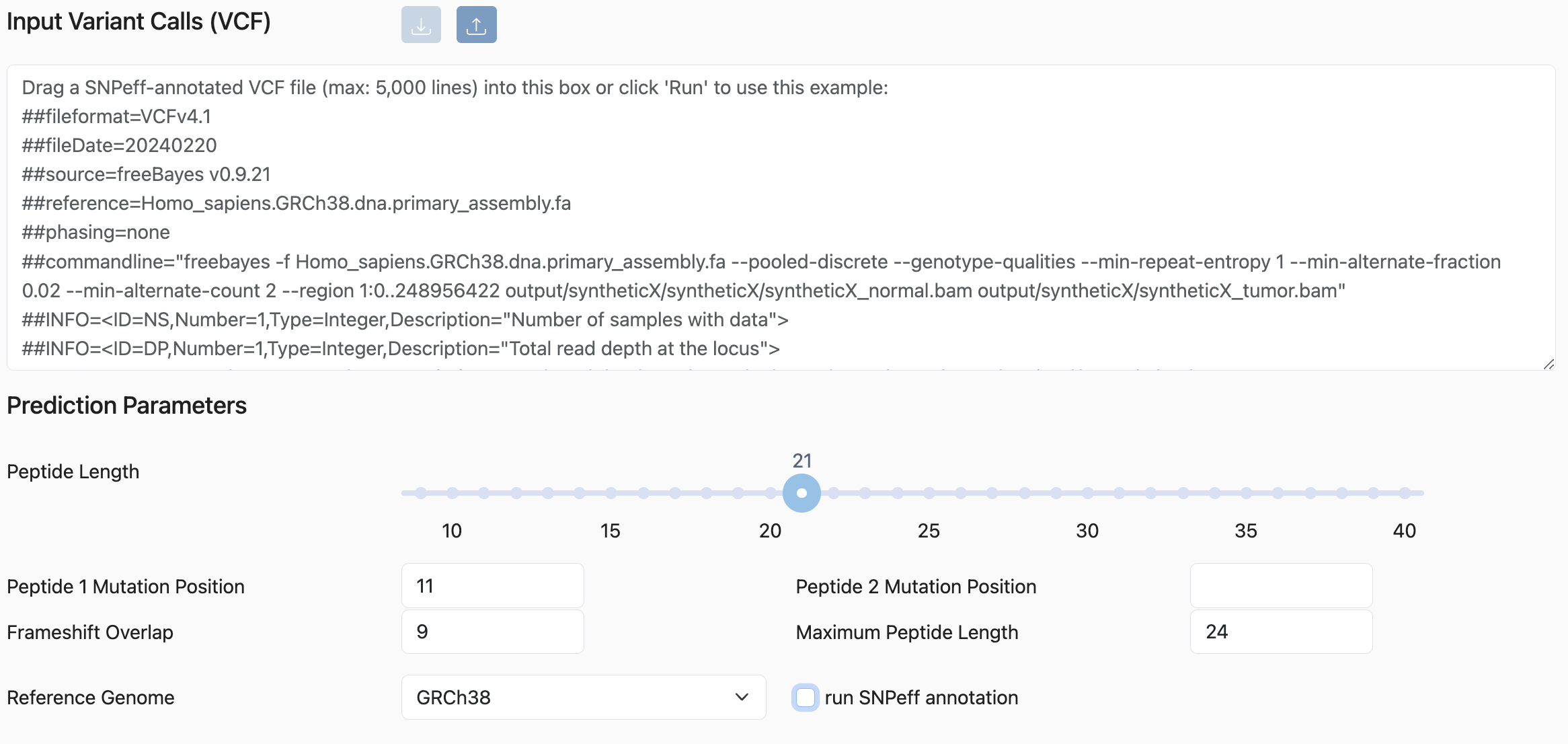
Peptide Length: The number of amino acids in the peptide.
Peptide 1 and 2 Mutation Position:
Position in the peptide where the mutation should be located. By default, only one peptide per variant is created. To create an additional peptide with the SNP at a different position, fill in the Peptide 2 Mutation Position field.
Frameshift Overlap:
Frameshift mutations often result in a long stretch of amino acids different from the reference. This parameter specifies the overlap length when breaking that stretch into overlapping peptides.
Maximum Peptide Length: Allows peptides longer than Peptide Length when needed (e.g., in-frame insertions, frameshifts, or variants near termini). Allowed values: Peptide Length + 1 to + 10 amino acids.
Reference Genome: Options include GRCh38 (default), GRCh37, and GRCm38/mm10. The selected genome should match the genome used to generate the VCF file.
run SNPeff annotation: If checked, SNPEff runs against the VCF before peptide generation. We recommend running annotation separately before uploading, as it can be time-consuming.
Results
Tabular Results
The following tables are available:
Variant Table: One row per variant (SNPs, MNPs, and Indels). Includes peptide pairs and warnings. Table controls: Download, Reset Table, Display Columns, Save Table State, and pagination.
Variant Table columns
peptide pairs: List of reference-mutant peptide pairs derived from this variant, along with peptide mutation position. Corresponding peptides will be found in the Peptide output table.
peptide warnings: Warnings from peptide generation for each variant. Includes warnings for successfully generated peptides and for variants where peptides could not be generated.
Peptide Table: One row per variant and affected transcript. Lists reference-mutant peptide pairs for each transcript.
Peptide Table columns
peptide pair id: Serial number for peptide pairs in the peptide-output table.
transcript reference allele: Reference allele (nucleotide) decoded from hgvs_dna.
transcript mutant allele: Tumor allele (nucleotide) decoded from hgvs_dna.
reference peptide: Reference peptide with requested PEPTIDELENGTH.
mutated peptide: Mutant peptide with requested PEPTIDELENGTH.
peptide mutation position: Position of the mutation within the peptide.
strand: Transcript strand: 1 for sense and -1 for anti-sense strand.
warnings: Any warnings associated with peptide generation for each reference-mutant peptide pair.
Unique Peptide Table: One row per variant and unique peptide. Peptides produced by multiple transcripts are collapsed; a representative transcript is selected.
Unique Peptide Table columns
chr: Chromosome.
position: Chromosomal position of mutation.
reference nucleotide: Reference nucleotide.
mutated nucleotide: Mutant nucleotide.
mutation effect: Predicted mutation effect (e.g., missense_variant, frameshift_variant, inframe_insertion, inframe_deletion).
gene name: HGNC gene symbol.
Ensembl gene accession: Ensembl gene identifier.
Ensembl transcript accession: Ensembl transcript identifier.
reference aa: Reference amino acid.
mutated aa: Mutated amino acid.
protein position: Mutation position in protein.
variant id: Internal unique identifier assigned to each variant.
mutation impact: SNPEff-predicted variant impact (LOW/MODERATE/HIGH).
transcript biotype: Classification of the transcript type (e.g., protein_coding, IG_ and TR_ types, nonsense_mediated_decay, non_stop_decay, pseudogene).
transcript mutation code: Mutation in HGVS format (nucleotide level) with coordinates based on the transcript.
protein mutation code: Mutation in HGVS format (amino acid level) with coordinates based on the protein.
cdna position: Mutation position in cDNA.
cds position: Mutation position in CDS.
Interpreting Results
The MPG tool outputs three tables: Variant, Peptide, and Unique Peptide. Use the column definitions above to interpret each table.
Note
The Unique Peptide table collapses peptides produced by multiple transcripts; the Peptide table lists all variant-transcript combinations; the Variant table summarizes at the variant level with peptide pairs and warnings.